
Antibody |
DNA modifications |
|
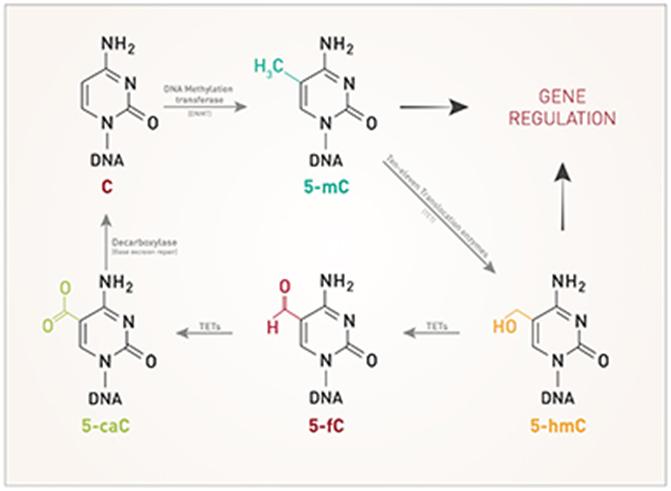
The pattern of DNA modifications is critical for genome stability and the control of gene expression in the cell. Methylation of 5-cytosine (5-mC), one of the best-studied epigenetic marks, is carried out by the DNA methyltransferases DNMT3A and B and DNMT1. DNMT3A and DNMT3B are responsible for de novo DNA methylation, whereas DNMT1 maintains existing methylation. 5-mC undergoes active demethylation which is performed by the Ten-Eleven Translocation (TET) familly of DNA hydroxylases. The latter consists of 3 members TET1, 2 and 3. All 3 members catalyze the conversion of 5-methylcytosine (5-mC) into 5-hydroxymethylcytosine (5-hmC), and further into 5-formylcytosine (5-fC) and 5-carboxycytosine (5-caC). 5-fC and 5-caC can be converted to unmodified cytosine by Thymine DNA Glycosylase (TDG). It is not yet clear if 5-hmC, 5-fC and 5-caC have specific functions or are simply intermediates in the demethylation of 5-mC. DNA methylation is generally considered as a repressive mark and is usually associated with gene silencing. It is essential that the balance between DNA methylation and demethylation is precisely maintained. Dysregulation of DNA methylation may lead to many different human diseases and is often observed in cancer cells. Diagenode offers highly validated antibodies against different proteins involved in DNA modifications as well as against the modified bases allowing the study of all steps and intermediates in the DNA methylation/demethylation pathway: |

|
Diagenode exclusively sources the original 5-methylcytosine monoclonal antibody (clone 33D3). Check out the list below to see all proposed antibodies for DNA modifications. |
|
Diagenode’s highly validated antibodies: ● Highly sensitive and specific ● Cost-effective (requires less antibody per reaction) ● Batch-specific data is available on the website ● Expert technical support ● Sample sizes available ● 100% satisfaction guarantee |

